Read news story coverage of our work:
https://www.me.gatech.edu/news/groundbreaking-lymphoma-tumor-model-paves-way-new-therapies
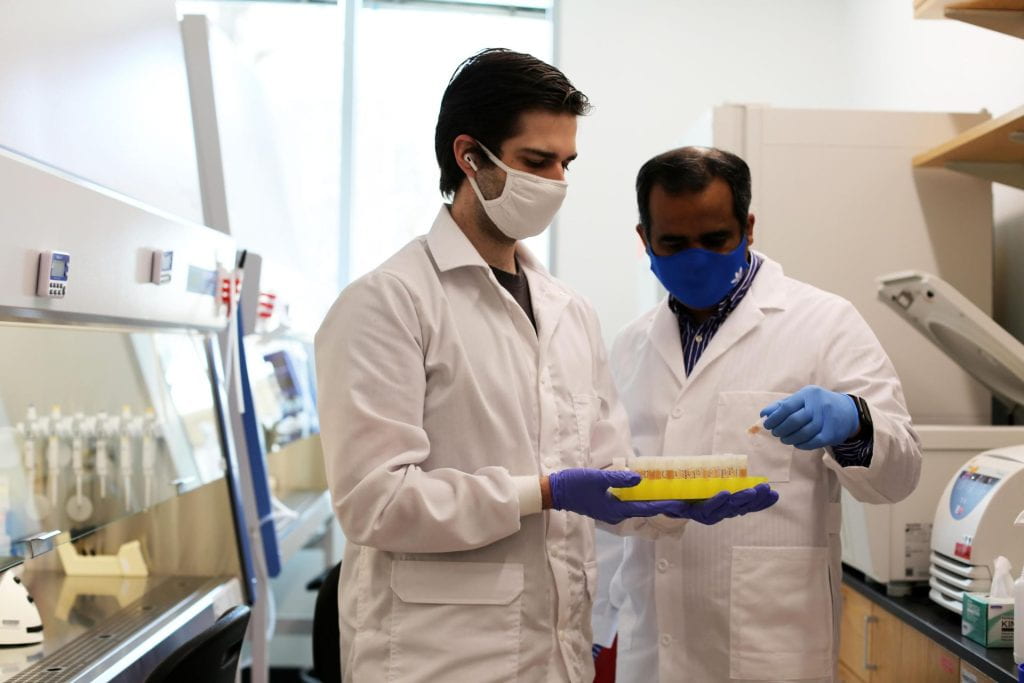

Professor Ankur Singh (right) with graduate student Chris Carlson, second author of the Nature Materials paper.
Groundbreaking Lymphoma Tumor Model Paves Way for New Therapies
March 29, 2023
By Catherine Barzler
In recent years, innovative cancer drugs that target specific molecular drivers of the disease have been embraced as the treatment of choice for many types of cancer. But despite significant advances, there is still a lack of understanding about how the complex interactions between a tumor and its surrounding environment in the body affect cancer progression. This problem has become a well-known roadblock in making novel treatments effective for more people.
Ankur Singh, professor in the George W. Woodruff School of Mechanical Engineering and the Wallace H. Coulter Department of Biomedical Engineering at Georgia Tech and Emory University led an international team of researchers in the development of a promising breakthrough for targeted cancer therapies.
The team bioengineered a synthetic tumor model to understand and then demonstrate how the tumor microenvironment impacts the effectiveness of targeted therapies for a specific type of lymphoma called Activated B Cell-like Diffuse Large B cell lymphoma (ABC-DLBCL). Their synthetic tumor model could change the game for designing and testing personalized cancer therapies. The research paper, which features an interdisciplinary team from institutions across the U.S. and around the world, was published in the journal Nature Materials.
A Cutting-Edge Tumor Model
Recent treatments for ABC-DLBCL that target specific molecular signals of the disease are in clinical trials. But, while the treatments have shown to be effective in lab testing (in vitro environments) and in mice (in vivo), they have proven less effective in humans, with over 60% of patients not responding.
“We wanted to understand how specific changes that happen in the microenvironment empower the lymphoma tumors to not respond to these drugs when administered in patients,” Singh said. “The ultimate goal is to build a patient-derived tissue model that represents the tumor and can be grown outside of the body, in order to truly understand the factors and conditions that control tumor behavior.”
To accurately test new therapies, a model tumor microenvironment should closely mimic the nuanced interactions that happen in a live tumor. But to understand those conditions, which can vary wildly from case to case, the researchers needed real patient data.
The researchers examined more than 1,100 ABC-DLBCL lymphoma patient samples to understand the molecular profiles of their tumors. For each sample, they used RNA sequencing and imaging to identify the composition, stiffness, and mechanical properties of the tumor tissue, along with other factors that play a role in how tumors grow and respond to treatment.
Combining what they learned from the patient data, the researchers designed a synthetic hydrogel-based model of the lymphoma tumor microenvironment. They bioengineered the model to have the specific qualities and characteristics seen in the microenvironments of the samples. Specifically, by modifying the hydrogel with cell-binding adhesive peptides and presenting immunological signals, they were able to recreate the intricate biological, chemical, and physical characteristics that are present in a live tumor microenvironment, including protein signals, tumor stiffness, and more. The customizable hydrogel proved to be supportive of tumor samples obtained from patients, a phenomenon that has not been previously demonstrated for lymphomas.
Combining Therapeutics
The team illustrated the viability of their model by testing how the tumors responded to a new type of inhibitor drug known as mucosa-associated lymphoid tissue lymphoma translocation protein 1 (MALT1 inhibitors) currently in human trials.
The researchers observed that, when being treated by MALT1, several tumor microenvironment factors related to the tumor cells — including T cell signal CD40 Ligand, collagen-like extracellular matrix, and the level of tissue stiffness — all empowered the tumor, helping the cancer cells resist responding to the new inhibitors even at high doses.
The researchers then sought a way to overcome the dampened tumor response by combining therapeutics that simultaneously suppress multiple aberrant oncogenic pathways in the same tumor cell. They found that when they used MALT1 and another inhibitor to target multiple pathways at the same time, they were able to promote more tumor death in the cells.
One of the major challenges is that tumors can engage multiple pathways in the cells to keep fueling the survival of tumor cells. However, the combination treatment was so powerful that, even in the presence of tumor microenvironment factors that supported tumor survival, they could still be overcome by the combination of therapies.
To validate the results from synthetic tissues developed in lab, the researchers then implanted actual patient tumors in an immunocompromised mouse model to determine how the patient tumors responded to the new therapies.
“We live in a world where we can claim a lot based on in vitro treatments, but the obvious question is always what happens in vivo,” Singh said. “What’s amazing is that we predicted this exact result in our synthetic model.”
Moving Forward
The researchers’ work clarifies the complex relationship between malignant ABC-DLBCL tumors and their dynamic surrounding environment, while highlighting the crucial importance of considering the tumor microenvironment when creating treatments that combine therapeutics.
The team’s work will help clinicians prioritize clinical trials of certain therapies and enable scientists to create more rational therapy combinations that could improve patient response rates to treatment. This is especially relevant for the potential of personalized treatment for lymphoma, as two individuals with the same cancer may benefit from different combinations and dosages of therapeutics.
A large portion of the patient samples used in creating the tumor models were provided by Emory through a collaboration with oncologist Jean Koff, one of the authors of the study.
“From a clinician’s standpoint, this work is very exciting because it exemplifies how findings from large genomic datasets may be translated into development of therapeutic strategies in lymphoma,” Koff said. “Singh’s cutting-edge organoid technology allows us to explore how patient-specific changes in the tumor microenvironment impact response to therapeutic agents, thus helping to deliver on the promise of precision medicine.”
The project further initiated a new partnership between teams at Georgia Tech and Emory and strengthened existing collaborations with Cornell Medicine. The teams will continue to work together to investigate molecular pathways that may be targeted to improve treatment outcomes for lymphoma patients.
The research comes at an important time in the field of drug testing. The FDA has begun to encourage alternatives to animal testing for pharmaceuticals. Singh’s powerful synthetic model that faithfully mimics real tumor environments is likely to be an example for other cancer researchers to follow for in vitro drug testing.
The research was funded by the National Institutes of Health, the National Cancer Institute, and the Wellcome Leap HOPE program.
Citation: Shah, S.B., Carlson, C.R., Lai, K. et al. Combinatorial treatment rescues tumour-microenvironment-mediated attenuation of MALT1 inhibitors in B-cell lymphomas. Nat. Mater. (2023).
DOI: https://doi.org/10.1038/s41563-023-01495-3

Ankur Singh, professor in the George W. Woodruff School of Mechanical Engineering and the Wallace H. Coulter Department of Biomedical Engineering at Georgia Tech and Emory University.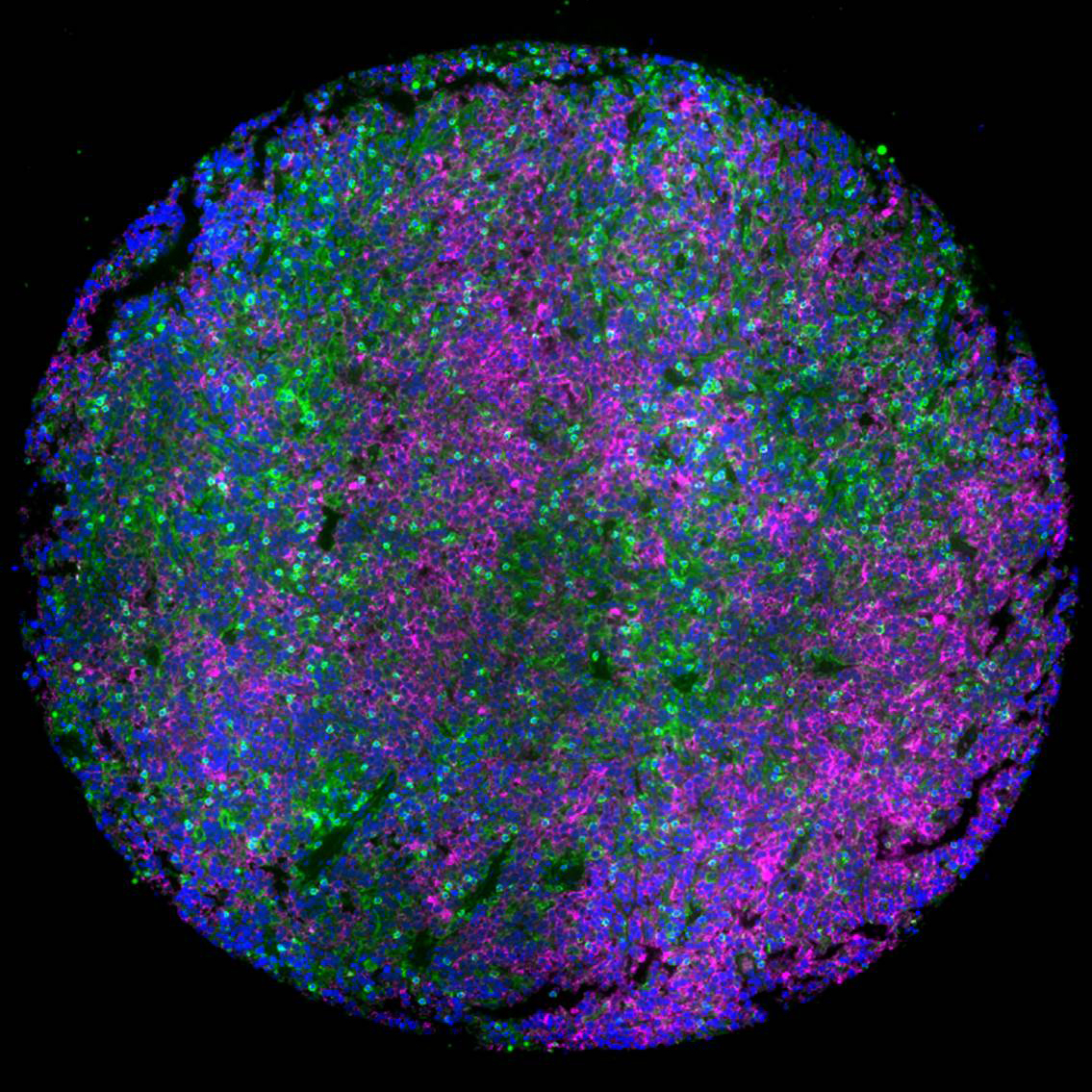
Lymph node tumor microenvironment of lymphomas with malignant B cells (magenta) surrounded by T cells (green), which induce survival mechanism in tumor cells and dampen the response to therapeutics. This microenvironment was faithfully mimicked in a 3D hydrogel of lymphoma patient tumors to discover role of tumor microenvironment in lymphomas and how combinatorial therapies can overcome resistance in tumors. Blue: Nucleus. Credit: Mayar Allam, Ahmet Coskun, and Ankur Singh (Georgia Tech)
Researchers pipetting organoids in Singh’s lab.